| RM ID | human_psi_associatedSNP_1 |
|---|---|
| Modification Type | PSI |
| Species | Human |
| Chromosome | chr1 |
| Position | 45285624 |
| Strand | (-) |
Reference Sequence  |
ATAAATCTTTCGCCTTTTACTAAAGATTTCCGTGGAGAGAA
|
Alterative Sequence  |
ATAAATCTTTCGCCTTTTACCAAAGATTTCCGTGGAGAGAA
|
| Gene | PTCH2 |
Gene Type  |
protein_coding |
| Gene Region | 3' UTR |
| Gene ID (Ensembl or others) | ENSG00000117425 |
Modification Status  |
human psi-loss variant |
Confidence Level  |
high |
Association Level  |
1 |
| Supported Studies | NA |
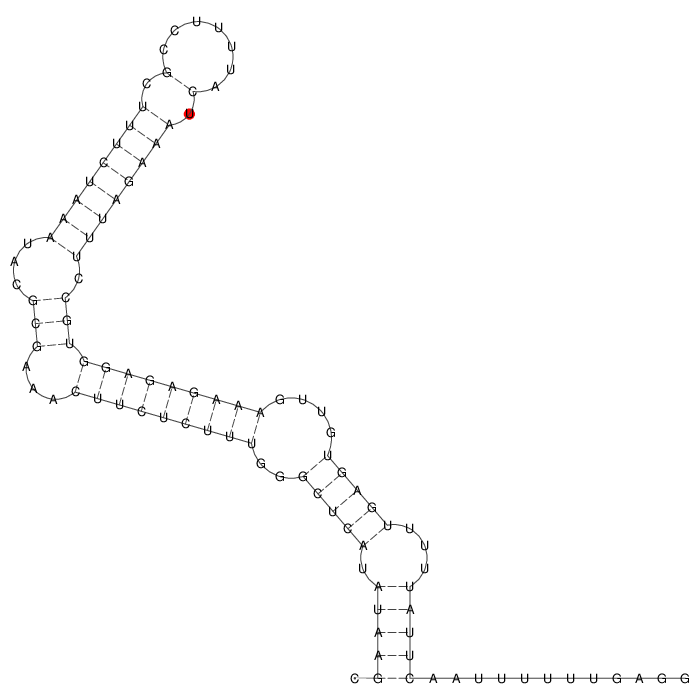
Predicted RNA Secondary Structure (Minimum Free Energy Fold by RNAFold)  |
.(((((.(((((..(((((((((...(((...((((((((.(.......).)))))))).))))))))))))....)))))...)))))............ (-23.90) |
Secondary Structure Graphical Visulization  |
|
| Genome Visualization |
| RM ID | human_psi_associatedSNP_1 |
|---|---|
| Variant Position | chr1:45285624 |
| Variant Source | dbSNP (v151) |
| Ref Base | A |
| Alt Base | G |
| TCGA Project | - |
| Barcode | - |
| RS ID | rs115827181 |
| dbSNP | View |
| Mut Region | 3' UTR |
| Mut Tpye | - |
| Deleterious Level | 0 |
| View this mutation in UCSC genome browser | View |
| View this mutation in Ensembl genome browser | View |
| RBP | Database | Study | Binding Region |
|---|---|---|---|
| SRRM4 | POSTAR2 | GSE57278 GSM1378374 |
chr1:45285540-45285640 |
| UPF1 | POSTAR2 | GSE69586 GSM1704214 |
chr1:45285540-45285660 |
| FIP1L1 | POSTAR2 | GSE37401 GSM917675 |
chr1:45285540-45285660 |
| PTBP1 | POSTAR2 | GSE57278 GSM1378377 |
chr1:45285540-45285660 |
| CPSF6 | POSTAR2 | GSE37401 GSM917665 |
chr1:45285553-45285627 |
| CPSF7 | POSTAR2 | GSE37401 GSM917663 |
chr1:45285555-45285664 |
| CSTF2T | POSTAR2 | GSE37401 GSM917677 |
chr1:45285559-45285663 |
| CSTF2 | POSTAR2 | GSE37401 GSM917676 |
chr1:45285561-45285629 |
| CPSF1 | POSTAR2 | GSE37401 GSM917672 |
chr1:45285566-45285626 |
| TARDBP | POSTAR2 | DRA001158 DRS012385 |
chr1:45285589-45285634 |
| METTL14 | POSTAR2 | GSE46705 GSM1135015 |
chr1:45285599-45285645 |
| CPSF4 | POSTAR2 | GSE37401 GSM917667 |
chr1:45285601-45285625 |
| FUS | POSTAR2 | DRA001158 DRS012384 |
chr1:45285601-45285626 |
| LIN28A | POSTAR2 | GSE44616 GSM1087848 |
chr1:45285601-45285630 |
| ATXN2 | POSTAR2 | DRA001158 DRS012389 |
chr1:45285601-45285643 |
| METTL3 | POSTAR2 | GSE46705 GSM1135006 |
chr1:45285601-45285645 |
| WTAP | POSTAR2 | GSE46705 GSM1135011 |
chr1:45285602-45285645 |
| TIA1 | POSTAR2 | E-MTAB-432 ERR039776-ERR039778 |
chr1:45285613-45285634 |
| HNRNPC | POSTAR2 | E-MTAB-1371 ERR196176-ERR196177-ERR196178-ERR196181-ERR196182-ERR196185-ERR196186-ERR196188-ERR196189 |
chr1:45285617-45285637 |
| TIAL1 | POSTAR2 | E-MTAB-432 ERR039779-ERR039781-ERR039782 |
chr1:45285617-45285638 |
| DIS3L2 | POSTAR2 | GSE81537 GSM2155555 |
chr1:45285620-45285640 |
| FBL | POSTAR2 | GSE43666 GSM1067864 |
chr1:45285620-45285640 |
| HNRNPA1 | POSTAR2 | GSE34996 GSM859978-GSM859979-GSM859980-GSM859981 |
chr1:45285620-45285640 |
| NOP56 | POSTAR2 | GSE43666 GSM1067863 |
chr1:45285620-45285640 |
| NOP58 | POSTAR2 | GSE43666 GSM1067861 |
chr1:45285620-45285640 |
| NUDT21 | POSTAR2 | GSE37401 GSM917661 |
chr1:45285620-45285640 |
| PRPF8 | POSTAR2 | ENCODE |
chr1:45285582-45285625 |
| miRNA Name | Target RNA | Target RNA Type | Source | Start Position | End Position |
|---|
| Gene Name | Genomic Location | Splicing Site | Relative Position |
|---|
| Variant | RS ID | Significant | Disease(s) | Clinvar Accession |
|---|
| Variant | RS ID | Pubmed ID | Disease | P value | Database | Type | TagSNP | Study_accession |
|---|